International Journal of Clinical and Medical Research | Year 2024 | Volume 1 | Issue 1 | Pages 39-45
A Review of Cholesterol Biosynthesis Pathway
Shatha R. Zaidan 1*1Department of Microbiology, Alkindy College of Medicine, University of Baghdad, Baghdad, Iraq
View PDF Download XML Download DOI XML DOI: 10.66590/ijcmr2024010103
Abstract
Cholesterol biosynthesis is an essential, tightly controlled biochemical process in human physiology. It is an essential part of membrane fluidity, cellular homeostasis, and for synthesis of steroid hormone and bile acid. The review will address the fine balance of enzymatic and genetic modulators governing endogenous cholesterol synthesis and emphasis will be laid on the importance and regulation. Mevalonate is produced first upon acetyl-CoA conversion. It is regulated by HMGCR enzyme that is also referred to as the rate limiting enzyme. Cholesterol is then biosynthesized through consecutive enzymatic conversions, with rate limiting enzymes including squalene epoxidase (SQLE), squalene synthase (FDFT1) and 7-dehydrocholesterol reductase (DHCR7). These enzymes are rigorously regulated by sterol regulatory element-binding proteins (SREBPs) through a complicated feedback mechanism that maintains metabolic balance. However, due to the multiple reactions involved, any dysfunction or disruptions in the expression of these regulatory genes, a variety disorders may be developed, comprising hyperlipidemia, atherosclerosis, various malignancies, and rare inherited enzymopathies, for instance, Smith-Lemli-Opitz syndrome (SLOS). From the clinical standpoint, various therapies have emerged to modulate major elements of the pathway, such as statins, PCSK9 inhibitors, and drugs to modulate SREBP signaling. These remain key to the management of cholesterol disorders and the betterment of patient outcomes
INTRODUCTION
Cell structure, signaling and homeostasis is maintained by cholesterol which is a crucial lipid molecule. It is fundamentally involved in preserving membrane structure and stability, influencing fluidity and permeability and precursor incorporated in the intrinsic production of steroid hormones, bile acids and vitamin D [1,2]. Although cholesterol can be acquired from diet, most is generated endogenously through the mevalonate pathway, a well-conserved metabolic pathway found in most eukaryotes [3]. This pathway comprises a sequence of enzymatic reactions that transform Acetyl-CoA into cholesterol, with each step tightly regulated by distinct genes and their corresponding enzymes. Tight regulatory mechanisms govern the cholesterol biosynthesis pathway to ensure physiological balance where cholesterol levels are maintained within a narrow range. Key regulatory mechanisms include transcriptional control by Sterol Regulatory Element-Binding Proteins (SREBPs), post-translational modifications of enzymes and feedback inhibition by cholesterol and its derivatives. Defects in sterol synthesis are causative factors for a broad spectrum of diseases, particularly cardiovascular disorders, metabolic syndromes and cancer [4]. For example, elevated cholesterol levels due to overactive biosynthesis constitute a key contributing factor to atherosclerosis and coronary artery disease, while any abnormalities in cholesterol synthesis promotes developmental disorders namely Smith-Lemli-Opitz Syndrome (SLOS) [5].
Given the central impact of cholesterol biosynthesis in health and disease, understanding the genes and regulatory mechanisms involved in this pathway is of paramount importance. Recent advances in molecular biology and genomics have deepened our understanding of the functional roles of specific genes and enzymes, highlighting their potential for therapeutic exploitation. The primary rationale of the present comprehensive review is to critically evaluate the molecular mechanisms underlying cholesterol biosynthesis pathway, focusing on the key genes involved, their regulation and their implications in health and disease. By synthesizing current knowledge, this article seeks to highlight emerging therapeutic strategies and future research directions in this critical area of metabolism. Furthermore, studying cholesterol biosynthesis pathway attracted researchers due not to the vital function of cholesterol isn’t in the cell membrane, proliferation and differentiation only, but since it was noticed that there is an enhancement in cholesterol biosynthesis in tumor cells [2]. Tumor cells require energy for their growth and proliferation therefore, tumor cells acquire this energy from cholesterol metabolism through reprogramming the process of cholesterol synthesis.
Cholesterol De Novo Biosynthesis Pathway
Cholesterol biosynthesis manifests primarily in the liver and proceeds via a multi-step enzymatic pathway initiated by Acetyl-CoA. The catalytic reaction governing the rate-limiting step of this process is facilitated by the enzyme 3-hydroxy-3-methylglutaryl-CoA reductase (HMGCR), which transforms HMG-CoA into mevalonate. Subsequent steps involve the formation of isoprenoids, squalene and lanosterol, eventually leading to cholesterol synthesis. The pathway is energetically demanding and is tightly regulated at multiple levels to balance cholesterol demand and synthesis [6]. In addition, the brain contains a high concentration of cholesterol, making up roughly a quarter of the body's total cholesterol supply. Cholesterol concentration in the brain ranges from 15 to 30 mg per gram of tissue, markedly higher than in most peripheral tissues, where levels typically range from 2 to 3 mg per gram of tissue. The blood-brain barrier imposes a significant limitation on the transport of cholesterol from the circulatory system into the brain, thereby regulating cerebral cholesterol homeostasis. As a result, de novo the endogenous synthesis of cholesterol is vital in preserving neuronal functionality and ensuring the structural integrity of cellular membranes within the nervous system. Cholesterol is indispensable for key neurological processes, including synapse formation, myelination and neurotransmission. Within the brain, astrocytes serve as the principal site of cholesterol biosynthesis, facilitating its delivery to neurons via apolipoprotein-mediated transport mechanisms. Disruptions in cholesterol biosynthetic pathways contribute to the onset and trajectory of neurodegenerative disorders, among these are Alzheimer's disease and Huntington's disease, highlighting the critical impact of cholesterol in maintaining neuronal health and molecular operation, where altered cholesterol metabolism may contribute to neuronal dysfunction and cognitive decline. Understanding the mechanisms regulating cholesterol synthesis in the brain could provide new therapeutic avenues for neurological disorders.
Genetic Regulation of Cholesterol Biosynthesis
Multiple genes are fundamentally participating in modulating cholesterol biosynthesis, orchestrating enzymatic pathways that ensure lipid homeostasis and proper neuronal function:
- HMGCR: It encodes the enzyme responsible for the rate-limiting step in cholesterol biosynthesis and constitutes the primary molecular target of statin therapy, which functions to modulate lipid metabolism and reduce cholesterol levels
- SREBF1 and SREBF2 (Sterol Regulatory Element-Binding Proteins): Transcription factors play a crucial role in modulating the expression of HMGCR and other enzymes essential for lipid metabolism. Through synchronized regulation of cholesterol-related genes, these proteins fine-tune metabolic flux across biosynthesis and transport pathways, preserving the delicate balance required for normal cellular function
- SCAP (SREBP Cleavage-Activating Protein): Promotes the activation of SREBP as a result of cholesterol levels
- LDLR (Low-Density Lipoprotein Receptor): Regulates cholesterol uptake and synthesis through feedback mechanisms
- LSS (Lanosterol Synthase): Catalyzes the cyclization of squalene into lanosterol, a crucial step in cholesterol biosynthesis
- SQLE (Squalene Epoxidase): Converts squalene to 2,3-oxidosqualene, a key intermediate in sterol biosynthesis
- CYP51A1 (Lanosterol 14α-Demethylase): Involved in demethylation of lanosterol, a critical step in the conversion to cholesterol
The interplay between these genes ensures that cholesterol synthesis is dynamically adjusted to meet cellular needs while preventing excessive accumulation.
Cholesterol Biosynthesis Pathway
The cholesterol biosynthesis pathway can be categorized into six distinct stages, each characterized by specific enzymes and intermediate compounds. The following provides a detailed overview of these sequential steps:
Synthesis of Mevalonate
- Starting Point: Acetyl-CoA (derived from glucose, fatty acids, or amino acids)
- Two molecules of acetyl-CoA condense to form acetoacetyl-CoA, catalyzed by acetoacetyl-CoA thiolase
- Acetoacetyl-CoA reacts with another acetyl-CoA to form 3-hydroxy-3-methylglutaryl-CoA (HMG-CoA), catalyzed by HMG-CoA synthase
- HMG-CoA is reduced to mevalonate by HMG-CoA reductase, the rate-limiting enzyme of the pathway
- Regulation: HMG-CoA reductase is tightly regulated by cholesterol levels, insulin and statin drugs
Conversion of Mevalonate to Isoprenoids
- Mevalonate is phosphorylated twice by mevalonate kinase and phosphomevalonate kinase to form mevalonate-5-pyrophosphate
- Mevalonate-5-pyrophosphate is decarboxylated by mevalonate diphosphate decarboxylase to form Isopentenyl Pyrophosphate (IPP), a 5-carbon isoprenoid
- IPP is isomerized to Dimethylallyl Pyrophosphate (DMAPP) by isopentenyl pyrophosphate isomerase
- IPP and DMAPP are the building blocks for all isoprenoids, including cholesterol, dolichol and ubiquinone
Synthesis of Farnesyl Pyrophosphate (FPP)
- IPP and DMAPP condense to form Geranyl Pyrophosphate (GPP), a 10-carbon intermediate
- GPP reacts with another IPP to form Farnesyl Pyrophosphate (FPP), a 15-carbon isoprenoid, catalyzed by farnesyl pyrophosphate synthase
- FPP is a key intermediate that diverges into cholesterol synthesis or other pathways (e.g., synthesis of dolichol, ubiquinone, or prenylated proteins)
Formation of Squalene
- Two molecules of FPP condense to form squalene, a 30-carbon linear hydrocarbon, catalyzed by squalene synthase (FDFT1)
- Squalene synthase is a key regulatory point, as it commits FPP to cholesterol biosynthesis
Cyclization of Squalene to Lanosterol
- Squalene is oxidized to squalene epoxide by Squalene Epoxidase (SQLE)
- Squalene epoxide undergoes cyclization to form lanosterol, the first sterol intermediate, catalyzed by Lanosterol Synthase (LSS)
- Lanosterol marks the transition from linear isoprenoids to cyclic sterols
Conversion of Lanosterol to Cholesterol
- Lanosterol undergoes a series of modifications, including demethylation, isomerization and reduction, to form cholesterol
- Key enzymes in this stage include:
- NSDHL (NAD(P)-dependent steroid dehydrogenase-like): Removes methyl groups
- HSD17B7 (hydroxysteroid 17-beta dehydrogenase 7): Catalyzes isomerization
- DHCR24 (24-dehydrocholesterol reductase): Reduces double bonds
- DHCR7 (7-dehydrocholesterol reductase): Catalyzes the final step, converting 7-dehydrocholesterol to cholesterol
- Feedback inhibition by cholesterol ensures tight control of the pathway. Flow chart of these stages are shown in Figure 1
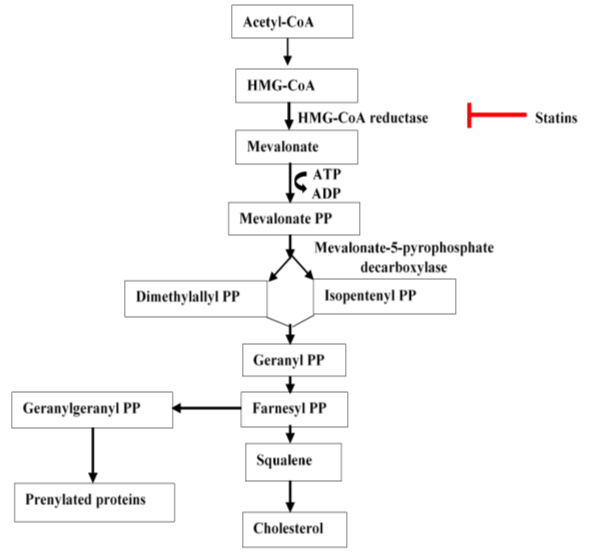
Figure 1: Cholesterol Synthesis in the Mevalonate Pathway [9]
The Role of SREBP Genes in Cholesterol Biosynthesis
A family of transcription factors known as "sterol regulatory element-binding proteins" (SREBPs) regulates the expression of genes necessary for the synthesis of fatty acids and cholesterol, hence playing a vital role in preserving lipid homeostasis. These transcription factors affect metabolic pathways through their regulatory role, maintaining cellular integrity and balanced lipid synthesis. Although the two main isoforms, SREBF1 and SREBF2, are encoded by different genetic loci, their control mechanisms share conserved regulatory characteristics. While SREBF2 is more specific to the biosynthesis of cholesterol, SREBF1 primarily affects the metabolism of fatty acids. The biogenesis of SREBPs involves synthesis of transcriptionally inactive precursor forms that are retained within the endoplasmic reticulum membrane which require cleavage by SCAP and S1P/S2P proteases to become active. They translocate to the nucleus after activation and attach to sterol regulatory elements (SREs) in the promoters of target genes to activate transcription. Translational dysregulation of SREBP is thought to be the main cause of some metabolic disorders, including hyperlipidemia, resistance of insulin and non-alcoholic fatty liver disease.
The SREBP family consists of three main isoforms encoded by two genes: SREBF1 and SREBF2. Below is a detailed overview of these genes and their roles:
SREBF1 (Sterol Regulatory Element-Binding Transcription Factor 1)
Function
- Encodes two major isoforms, SREBP-1a and SREBP-1c, through alternative splicing
- SREBP-1a: Regulates genes involved in both cholesterol and fatty acid synthesis
- SREBP-1c: Primarily controls fatty acid synthesis and lipogenesis
Regulation
- Activated by insulin and inhibited by Polyunsaturated Fatty Acids (PUFAs)
- Responds to nutrient availability, particularly carbohydrates
Target Genes
Includes FASN (Fatty Acid Synthase), ACC (Acetyl-CoA carboxylase) and SCD1 (stearoyl-CoA desaturase 1).
Clinical Relevance
Hyperactivation of Sterol Regulatory Element-Binding Protein 1 (SREBP-1) demonstrates a significant correlation with obesity phenotypes, insulin resistance and Non-Alcoholic Fatty Liver Disease (NAFLD) [7].
SREBF2 (Sterol Regulatory Element-Binding Transcription Factor 2)
Function
- Encodes SREBP-2, which primarily regulates cholesterol biosynthesis and uptake
- Specifically activates genes involved in the mevalonate pathway and LDL Receptor (LDLR) expression
Regulation
- Activated by low intracellular cholesterol levels
- Inhibited by cholesterol and oxysterols through feedback mechanisms
Target Genes
Includes HMGCR (HMG-CoA reductase), SQLE (squalene epoxidase) and LDLR (low-density lipoprotein receptor).
Regulation of SREBP Genes
Transcriptional Control
- SREBPs are synthesized as inactive precursors bound to the Endoplasmic Reticulum (ER) membrane
- Low sterol levels trigger their proteolytic cleavage and translocation to the nucleus, where they activate target gene transcription
Post-Translational Modifications
Phosphorylation, ubiquitination and acetylation modulate SREBP activity and stability.
Interplay with Other Pathways
SREBPs interact with insulin signaling, mTOR (mechanistic target of rapamycin) and AMPK (AMP-activated protein kinase) pathways to coordinate lipid metabolism with cellular energy status.
Disease Associations of Genetic Variations in Cholesterol Biosynthesis
Genetic variations in cholesterol biosynthesis pathways have been linked to several disorders:
- Familial Hypercholesterolemia (FH): Mutations in LDLR, PCSK9 and APOB lead to high cholesterol levels and premature cardiovascular disease
- Metabolic Syndrome: Dysregulation of cholesterol biosynthesis genes contributes to obesity, insulin resistance and hypertension
- Neurodegenerative Diseases: Abnormal cholesterol metabolism has been implicated in Alzheimer's disease and Huntington's disease.
- Metabolic Disorders: Over activation of SREBP-1 contributes to fatty liver disease, obesity and type 2 diabetes. Moreover, SREBP-2 dysregulation is implicated in hypercholesterolemia and atherosclerosis
- Cancer: SREBPs are upregulated in many cancers to support rapid cell proliferation by providing lipids for membrane synthesis and signaling molecules [8]
- Therapeutic Targets:
- Statins indirectly inhibit SREBP-2 by lowering cholesterol levels
- Emerging therapies target SREBP cleavage-activating protein (SCAP) or SREBP-specific inhibitors to modulate lipid metabolism in metabolic and oncologic diseases
Factors Affecting Gene Regulation in Cholesterol Biosynthesis
Several factors are affecting gene regulation in cholesterol biosynthesis, encompassing genetic, epigenetic and environmental influences. Transcriptional regulation occurs through essential elements, Sterol Regulatory Elements (SREs), which modulate gene expression in accordance with intracellular cholesterol concentrations. Furthermore, gene expression is highly modified by DNA methylation and histone acetylation (chromatin modification) which achieved as a result of altering nucleosome positioning and the interactions between DNA and proteins. In addition, non-coding RNAs, particularly microRNAs (miRNAs), contribute to post-transcriptional regulation by targeting mRNA transcripts of key enzymes such as HMGCR (3-hydroxy-3-methyl-glutaryl-CoA reductase) and SREBF2 (Sterol Regulatory Element Binding Protein 2). Moreover, external factors such as dietary intake, hormonal signals and pharmacological treatments can influence gene regulation, resulting in variations in cholesterol biosynthesis and overall metabolic health. A comprehensive understanding of these regulatory mechanisms is crucial for the development of targeted therapies aimed at managing disorders related to cholesterol metabolism.
Therapeutic Targets
Targeting cholesterol biosynthesis at the genetic and molecular level has led to several therapeutic approaches:
- Statins: Therapeutic targeting of HMGCR, the principal regulatory enzyme in the cholesterol synthesis pathway, effectively decreases plasma cholesterol levels through suppression of endogenous cholesterol production. Additionally, statins indirectly inhibit SREBP-2 by lowering cholesterol levels
- PCSK9 Inhibitors: Enhance LDL receptor recycling, promoting cholesterol clearance from circulation
- Ezetimibe: Reduces intestinal cholesterol absorption, lowering plasma cholesterol levels
- Gene Therapy Approaches: Emerging techniques targeting SREBF2, SCAP and INSIG genes to regulate cholesterol homeostasis
- Squalene Synthase Inhibitors: Target downstream cholesterol synthesis steps to reduce cholesterol levels without affecting other isoprenoid pathways
Future Directions
The future of cholesterol biosynthesis research lies in integrating cutting-edge technologies, such as single-cell omics, CRISPR and AI, with a deeper understanding of pathway regulation and its role in health and disease. Below are some key areas of focus that represent the next frontier in this field:
Single-Cell Omics and Spatial Transcriptomics
Single-Cell Analysis
- Investigate cholesterol biosynthesis at the single-cell level to understand cell-to-cell variability in cholesterol metabolism
- Identify rare cell populations with unique cholesterol biosynthesis profiles, such as cancer stem cells or immune cells
Spatial Transcriptomics
Map cholesterol biosynthesis gene expression within tissues to uncover spatial heterogeneity and microenvironmental influences.
Integration of Multi-Omics Data
- Holistic Approaches: Combine genomics, transcriptomics, proteomics, metabolomics and epigenomics to unravel the complex regulatory networks controlling cholesterol biosynthesis
- Network Modeling: Develop systems biology models to predict how perturbations in one molecular layer (e.g., genetic mutations) affect the entire pathway
Precision Medicine and Personalized Therapies
- Genetic Profiling: Use genomic data to identify individuals with mutations in cholesterol biosynthesis genes (e.g., DHCR7, HMGCR) and tailor treatments accordingly
- Pharmacogenomics: Study how genetic variations influence responses to cholesterol-lowering drugs like statins and PCSK9 inhibitors
- Biomarker Discovery: Identify multi-omics biomarkers for early diagnosis and monitoring of cholesterol-related disorders
Gene Editing and CRISPR Technology
Therapeutic Applications
- Use CRISPR-Cas9 to correct mutations in cholesterol biosynthesis genes (e.g., in Smith-Lemli-Opitz syndrome)
- Develop gene therapies to modulate the expression of key genes like HMGCR or SREBF2
Functional Studies
Perform high-throughput CRISPR screens to identify novel regulators of cholesterol biosynthesis.
Targeting SREBP and Pathway Regulation
- SREBP Inhibitors: Develop small molecules or biologics to inhibit SREBP activation and reduce cholesterol biosynthesis in cancer or metabolic diseases
- Post-Translational Modifications: Explore the role of phosphorylation, ubiquitination and acetylation in regulating SREBP and cholesterol biosynthesis enzymes
Role of Cholesterol Biosynthesis in Non-Hepatic Tissues
- Brain and Neurodegeneration: Elucidate the contribution of de novo cholesterol biosynthesis within the central nervous system to the pathogenesis of neurodegenerative disorders, particularly Alzheimer's disease and Parkinson's disease
- Immune System: Study how cholesterol biosynthesis in immune cells influences inflammation and autoimmune diseases
- Cancer Metabolism: Delineate the functional contribution of endogenous cholesterol biosynthesis pathways to neoplastic advancement and the development of therapeutic resistance in malignant cells
Artificial Intelligence and Machine Learning
- Data Analysis: Use AI to analyze large multi-omics datasets and identify novel patterns, biomarkers and therapeutic targets
- Predictive Modeling: Develop machine learning models to predict disease risk, progression and treatment outcomes based on cholesterol biosynthesis profiles
Environmental and Lifestyle Influences
- Nutrigenomics: Study how diet and lifestyle (e.g., high-fat diets, exercise) influence cholesterol biosynthesis gene expression
- Epigenetics: Investigate how environmental factors (e.g., toxins, stress) alter epigenetic regulation of cholesterol biosynthesis genes
Rare Genetic Disorders
- Smith-Lemli-Opitz Syndrome (SLOS): Develop targeted therapies for SLOS and other rare disorders caused by defects in cholesterol biosynthesis genes
- Newborn Screening: Implement screening programs to identify and treat cholesterol biosynthesis disorders early in life
Therapeutic Innovations
- Novel Drug Targets: Discover and experimentally verify novel enzymatic components and regulatory factors within the cholesterol biosynthesis cascade as potential therapeutic targets
- Nanotechnology: Develop nanoparticle-based delivery systems for targeted modulation of cholesterol biosynthesis in specific tissues
CONCLUSIONS
According to the researchers surveyed and results achieved by these studies, the followings can be concluded:
- The cholesterol biosynthesis pathway represents a cornerstone of cellular metabolism, with its constituent genes playing indispensable roles in maintaining physiological homeostasis and driving pathological processes. This review underscores the critical functions of key genes such as HMGCR, SQLE and DHCR7, which orchestrate the conversion of acetyl-CoA into cholesterol while balancing the production of essential non-sterol isoprenoids
- The biosynthetic pathway of cholesterol is stringently controlled to equilibrate cellular cholesterol concentrations within a narrow span. Major mechanisms of its regulation also involve transcriptional regulation by SREBPs, post-translational modification of enzymes and feedback inhibition by cholesterol and its oxysterols. Violation of this signalling cascade is associated with different pathologies, such as cardiovascular and metabolic diseases and cancer. These associations highlight a core biological principle: maintaining homeostatic balance is critical for preserving normal physiological function and preventing disease
- Independent but parallel research on gene-editing technologies like CRISPR-Cas9 and the burgeoning field of metabolomics—has broadened the scope of research into cholesterol metabolism. Together with emerging therapeutic strategies, such as the new class of cholesterol-lowering drugs (statins and PCSK9 inhibitors) and interventions that target SREBP signaling, they offer unprecedented opportunities to explore tissue-specific aspects of cholesterol metabolism. This, in turn, is leading to the emergence of personalized therapies that take advantage of an individual's unique genetic and metabolic landscape
- Notwithstanding significant advances in elucidating the cholesterol biosynthetic pathway, important gaps remain—particularly concerning how regulation varies across different tissues and the role of non-coding RNAs in controlling key metabolic genes. This less-explored areas were considered in this review by examining the dynamic interplay between genetic regulatory networks and environmental factors that influence cholesterol metabolism. Deciphering these intricate relationships could identify new therapeutic targets to facilitate the design of more effective and targeted pharmacological treatments
- The importance of CRISPR-based gene-editing technologies has attracted growing interest in targeting pivotal regulatory genes, such as HMGCR and SREBP, to develop precision therapies for dyslipidemias. This review offers a focused exploration of these promising therapeutic strategies, addressing a topic that remains relatively underrepresented in current literature. The intersection of molecular genetics, innovative biotechnologies and lipid metabolic regulation presents significant opportunities for revolutionizing therapeutic approaches to cholesterol disorders through genetic interventions
- Given the complex nature of cholesterol homeostasis and its involvement in various diseases, further research is needed to explore the genetic determinants of inter-individual variability in cholesterol metabolism. In particular, the application of multi-omics approaches (such as metabolomics and genomics) could provide a more holistic view of how genetic regulation interacts with environmental factors in disease progression
Ethical Statement
The project was approved by the local ethical committee at University of Baghdad -Alkindy college of Medicine.
I hereby confirm that all the Figure in the manuscript is cited. Furthermore, any Figures and images, that are not mine/ours, have been included with the necessary permission for re-publication, which is attached to the manuscript.
REFERENCES
- Luo, J. et al. “Mechanisms and regulation of cholesterol homeostasis.” Nat Rev Mol Cell Biol, vol. 21, no. 4, 2020, pp. 225–245. https://doi.org/10.1038/s41580-019-0190-7.
- Luo, J. et al. “Cholesterol metabolism in cancer: Mechanisms and therapeutic opportunities.” Nat Metab, vol. 2, no. 2, 2020, pp. 132–141. https://doi.org/10.1038/s41591-019-0731-2.
- Goldstein, J.L. et al. “A century of cholesterol and coronaries: From plaques to genes to statins.” Cell, vol. 161, no. 1, 2015, pp. 161–172. https://doi.org/10.1016/j.cell.2015.01.036.
- Porter, F.D. et al. “Malformation syndromes caused by disorders of cholesterol synthesis.” J Lipid Res, vol. 52, no. 1, 2011, pp. 6–34. https://doi.org/10.1194/jlr.R009548.
- Waterham, H.R. et al. “Mutations in the 3β-Hydroxysterol Δ24-Reductase gene cause desmosterolosis, an autosomal recessive disorder of cholesterol biosynthesis.” Am J Hum Genet, vol. 69, no. 4, 2001, pp. 685–694. https://doi.org/10.1086/323473.
- Zerenturk, E.J. et al. “Desmosterol and DHCR24: unexpected new directions for a terminal step in cholesterol synthesis.” Prog Lipid Res, vol. 52, no. 4, 2013, pp. 666–680. https://doi.org/10.1016/j.plipres.2013.09.002.
- Shimano, H. et al. “SREBP-regulated lipid metabolism: Convergent physiology—divergent pathophysiology.” Nat Rev Endocrinol, vol. 13, no. 12, 2017, pp. 710–730. https://doi.org/10.1038/nrendo.2017.91.
- Cheng, C. et al. “Lipid metabolism reprogramming and its potential targets in cancer.” Cancer Commun, vol. 41, no. 5, 2021, pp. 371–388. https://doi.org/10.1002/cac2.12144.
- Zaidan, R.S. Effect of Obesity on Blood Brain Barrier Integrity in Ischemic Brain Using Mouse Model. University of Saarland, 2020. Dissertation.

